Potassium »
PDB 3ay9-3ccl »
3b7i »
Potassium in PDB 3b7i: Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid
Enzymatic activity of Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid
All present enzymatic activity of Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid:
3.4.11.10;
3.4.11.10;
Protein crystallography data
The structure of Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid, PDB code: 3b7i
was solved by
N.J.Ataie,
Q.Q.Hoang,
M.P.D.Zahniser,
A.Milne,
G.A.Petsko,
D.Ringe,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 20.37 / 1.75 |
| Space group | P 61 2 2 |
| Cell size a, b, c (Å), α, β, γ (°) | 109.907, 109.907, 91.095, 90.00, 90.00, 120.00 |
| R / Rfree (%) | 17.7 / 20.9 |
Other elements in 3b7i:
The structure of Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid also contains other interesting chemical elements:
| Zinc | (Zn) | 2 atoms |
| Sodium | (Na) | 2 atoms |
Potassium Binding Sites:
The binding sites of Potassium atom in the Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid
(pdb code 3b7i). This binding sites where shown within
5.0 Angstroms radius around Potassium atom.
In total only one binding site of Potassium was determined in the Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid, PDB code: 3b7i:
In total only one binding site of Potassium was determined in the Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid, PDB code: 3b7i:
Potassium binding site 1 out of 1 in 3b7i
Go back to
Potassium binding site 1 out
of 1 in the Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid
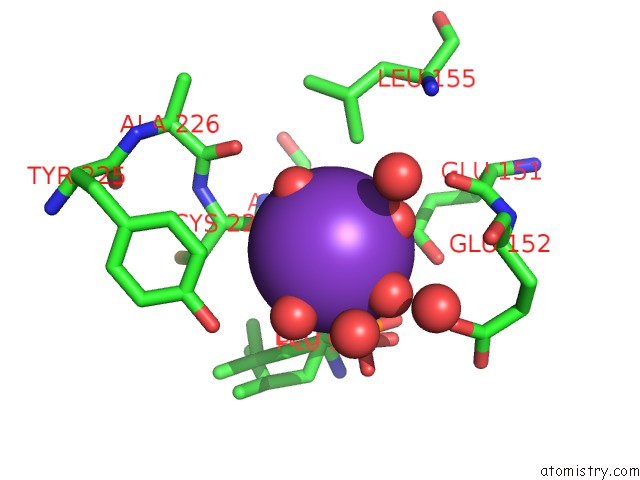
Mono view
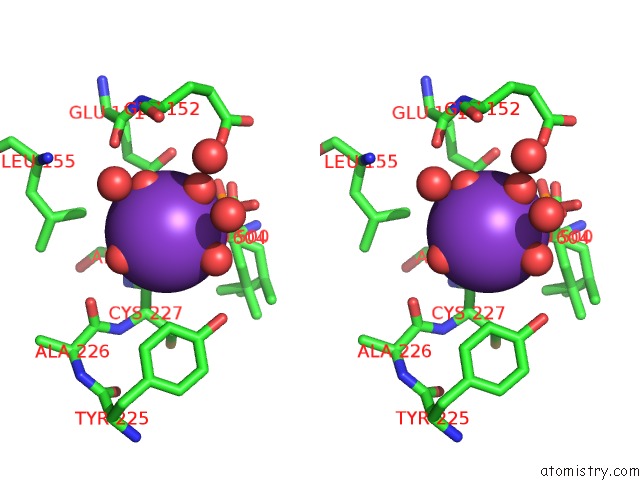
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 1 of Crystal Structure of the S228A Mutant of the Aminopeptidase From Vibrio Proteolyticus in Complex with Leucine Phosphonic Acid within 5.0Å range:
|
Reference:
N.J.Ataie,
Q.Q.Hoang,
M.P.Zahniser,
Y.Tu,
A.Milne,
G.A.Petsko,
D.Ringe.
Zinc Coordination Geometry and Ligand Binding Affinity: the Structural and Kinetic Analysis of the Second-Shell Serine 228 Residue and the Methionine 180 Residue of the Aminopeptidase From Vibrio Proteolyticus. Biochemistry V. 47 7673 2008.
ISSN: ISSN 0006-2960
PubMed: 18576673
DOI: 10.1021/BI702188E
Page generated: Sat Aug 9 04:32:23 2025
ISSN: ISSN 0006-2960
PubMed: 18576673
DOI: 10.1021/BI702188E
Last articles
K in 5PA3K in 5S9L
K in 5P9D
K in 5P9C
K in 5P9A
K in 5OX7
K in 5OWO
K in 5OU5
K in 5OSN
K in 5OSX