Potassium »
PDB 7zri-8bm0 »
8b2s »
Potassium in PDB 8b2s: GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain
Protein crystallography data
The structure of GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain, PDB code: 8b2s
was solved by
O.V.Moroz,
E.Blagova,
A.A.Lebedev,
L.K.Skov,
R.A.Pache,
K.M.Schnorr,
L.Kiemer,
S.Nymand-Grarup,
L.Ming,
L.Ye,
M.Klausen,
M.T.Cohn,
E.G.W.Schmidt,
G.J.Davies,
K.S.Wilson,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 46.58 / 1.94 |
| Space group | P 31 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 99.438, 99.438, 133.254, 90, 90, 120 |
| R / Rfree (%) | 16.8 / 20.1 |
Potassium Binding Sites:
The binding sites of Potassium atom in the GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain
(pdb code 8b2s). This binding sites where shown within
5.0 Angstroms radius around Potassium atom.
In total 2 binding sites of Potassium where determined in the GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain, PDB code: 8b2s:
Jump to Potassium binding site number: 1; 2;
In total 2 binding sites of Potassium where determined in the GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain, PDB code: 8b2s:
Jump to Potassium binding site number: 1; 2;
Potassium binding site 1 out of 2 in 8b2s
Go back to
Potassium binding site 1 out
of 2 in the GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain
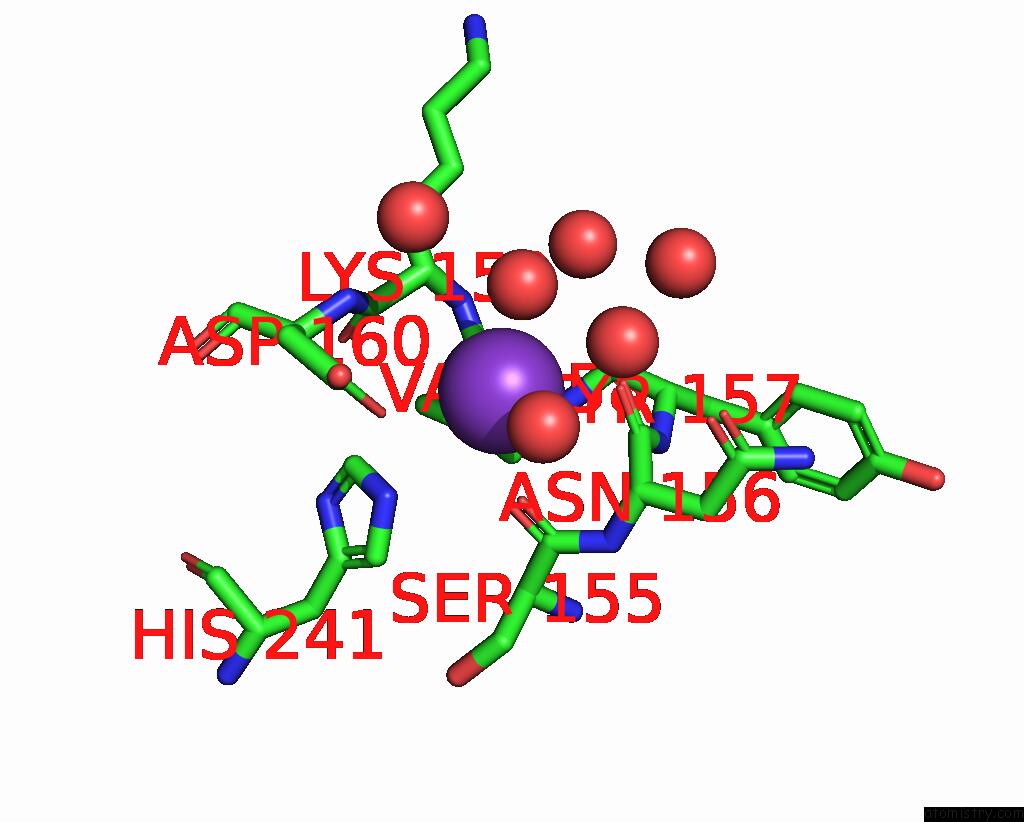
Mono view
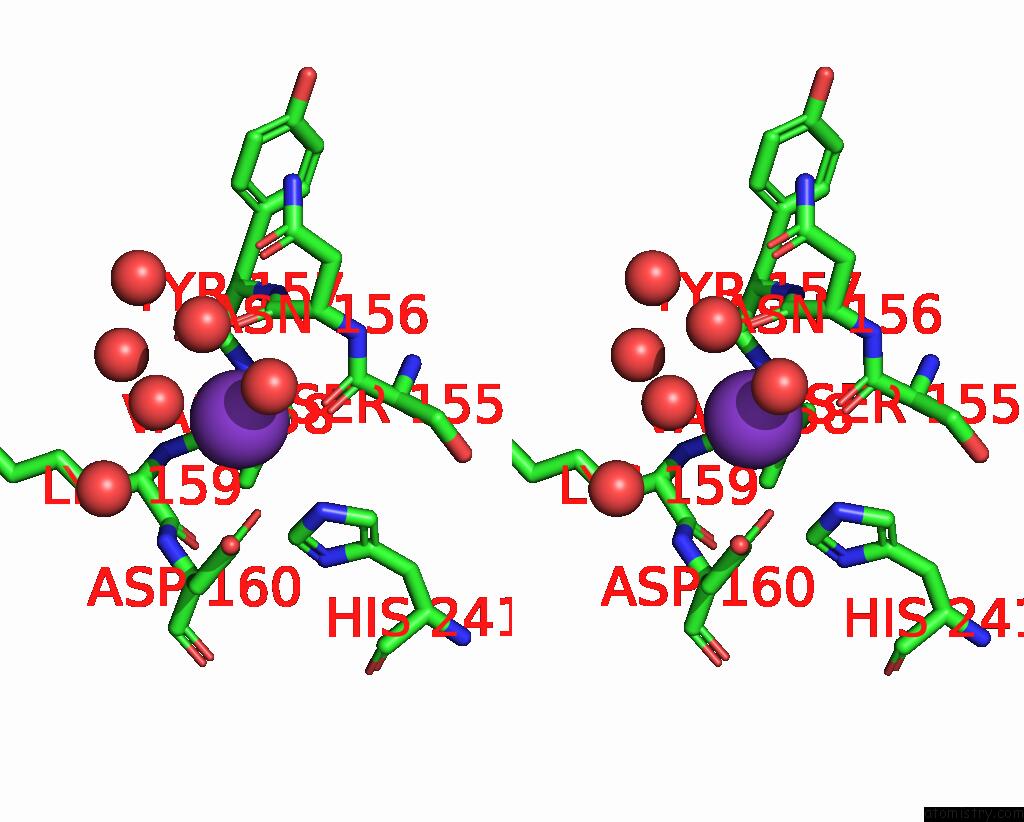
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 1 of GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain within 5.0Å range:
|
Potassium binding site 2 out of 2 in 8b2s
Go back to
Potassium binding site 2 out
of 2 in the GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain
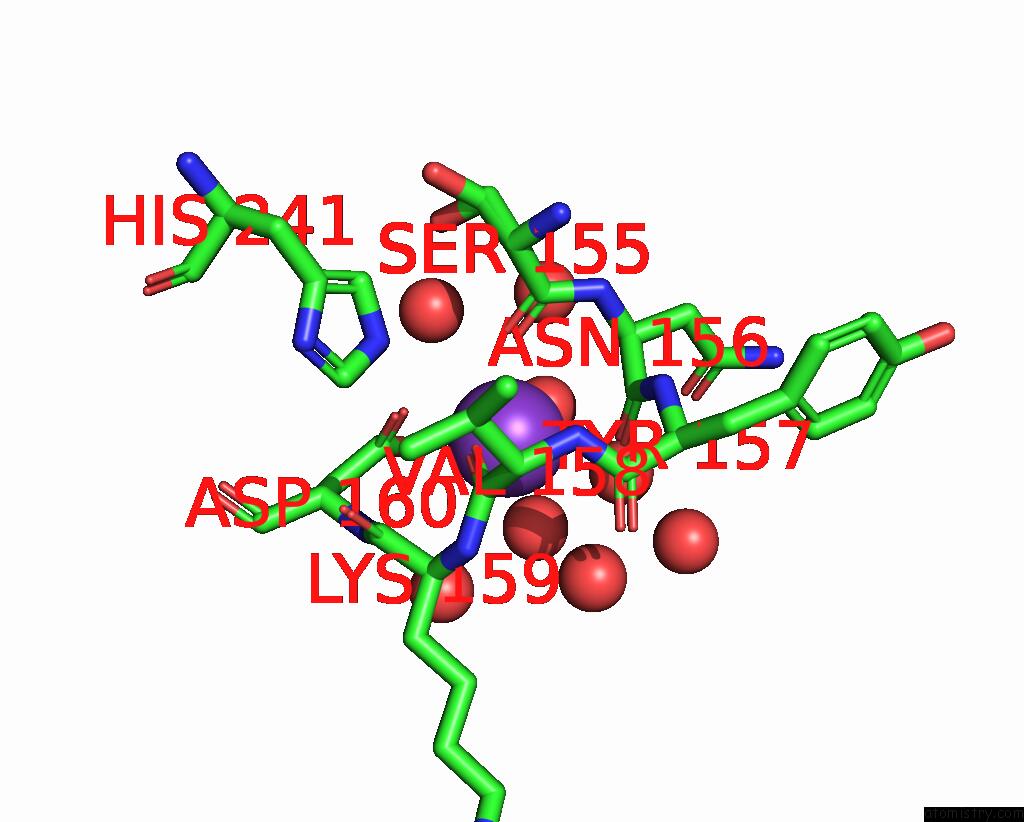
Mono view
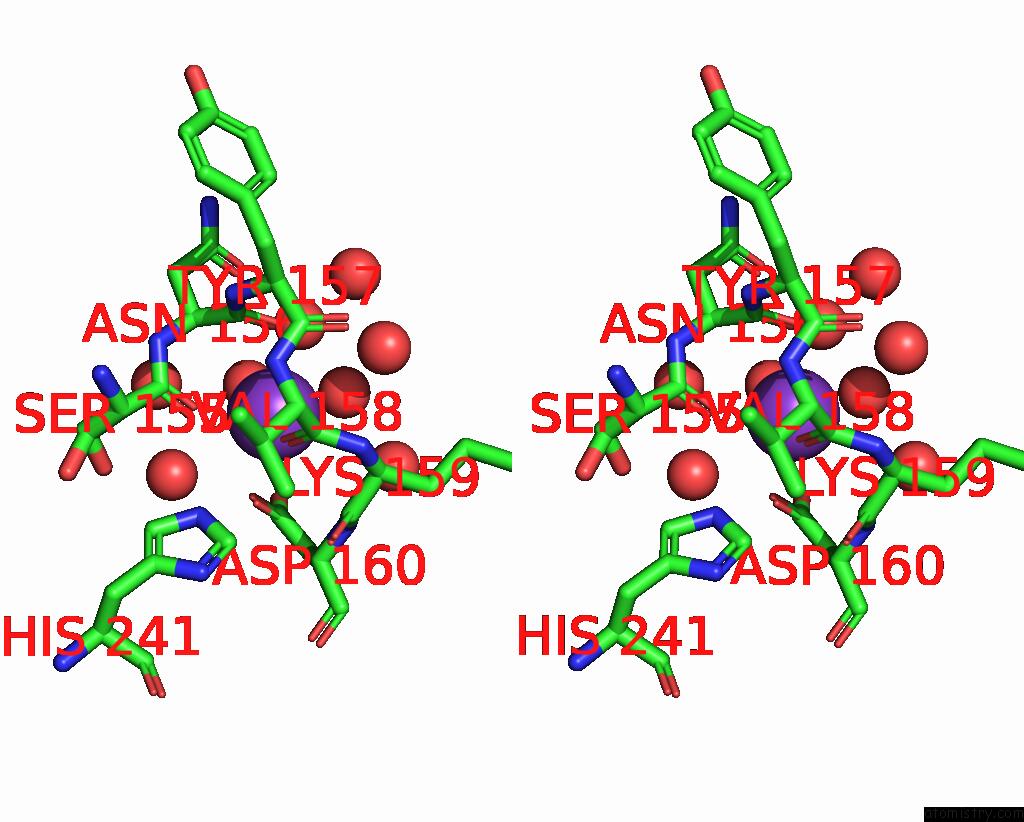
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 2 of GH24 Family Muramidase From Trichophaea Saccata with An SH3-Like Cell Wall Binding Domain within 5.0Å range:
|
Reference:
O.V.Moroz,
E.Blagova,
A.A.Lebedev,
L.K.Skov,
R.A.Pache,
K.M.Schnorr,
L.Kiemer,
E.P.Friis,
S.Nymand-Grarup,
L.Ming,
L.Ye,
M.Klausen,
M.T.Cohn,
E.G.W.Schmidt,
G.J.Davies,
K.S.Wilson.
Module Walking Using An SH3-Like Cell-Wall-Binding Domain Leads to A New GH184 Family of Muramidases. Acta Crystallogr D Struct 2023BIOL.
ISSN: ISSN 2059-7983
PubMed: 37428847
DOI: 10.1107/S2059798323005004
Page generated: Mon Aug 12 22:01:43 2024
ISSN: ISSN 2059-7983
PubMed: 37428847
DOI: 10.1107/S2059798323005004
Last articles
Zn in 9JPJZn in 9JP7
Zn in 9JPK
Zn in 9JPL
Zn in 9GN6
Zn in 9GN7
Zn in 9GKU
Zn in 9GKW
Zn in 9GKX
Zn in 9GL0