Potassium »
PDB 6x90-6z2v »
6z2v »
Potassium in PDB 6z2v: CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3)
Enzymatic activity of CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3)
All present enzymatic activity of CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3):
2.7.12.1;
2.7.12.1;
Protein crystallography data
The structure of CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3), PDB code: 6z2v
was solved by
M.Schroeder,
A.Chaikuad,
F.Bracher,
S.Knapp,
Structural Genomicsconsortium (Sgc),
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 39.37 / 2.60 |
| Space group | I 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 84.376, 45.393, 106.334, 90.00, 111.16, 90.00 |
| R / Rfree (%) | 20.2 / 26.5 |
Other elements in 6z2v:
The structure of CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3) also contains other interesting chemical elements:
| Chlorine | (Cl) | 2 atoms |
Potassium Binding Sites:
The binding sites of Potassium atom in the CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3)
(pdb code 6z2v). This binding sites where shown within
5.0 Angstroms radius around Potassium atom.
In total only one binding site of Potassium was determined in the CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3), PDB code: 6z2v:
In total only one binding site of Potassium was determined in the CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3), PDB code: 6z2v:
Potassium binding site 1 out of 1 in 6z2v
Go back to
Potassium binding site 1 out
of 1 in the CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3)
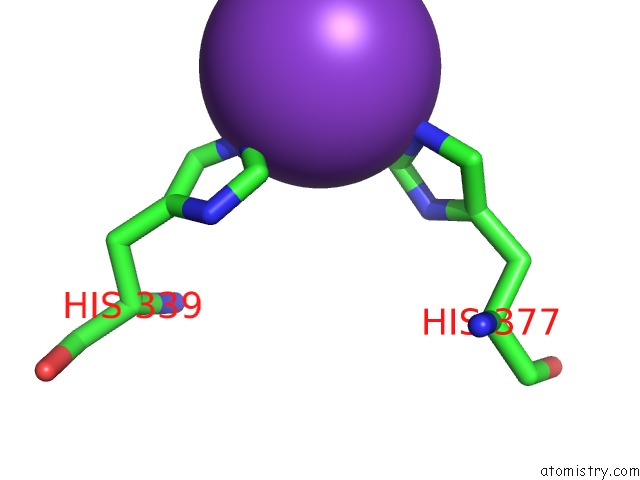
Mono view
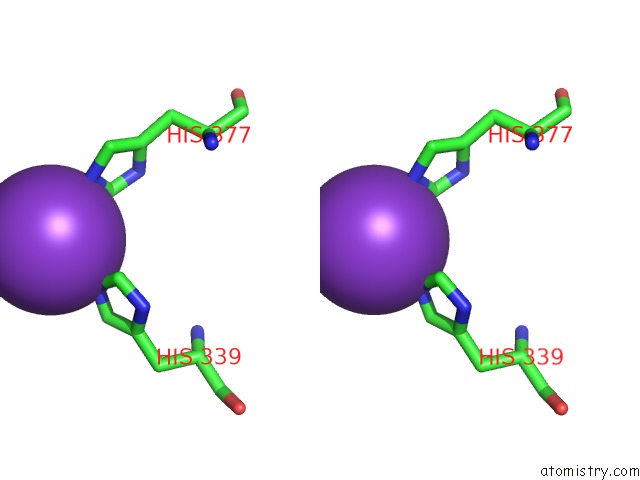
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 1 of CLK3 A319V Mutant Bound with Beta-Carboline Kh-CARB13 (Cpd 3) within 5.0Å range:
|
Reference:
M.Schroder,
A.N.Bullock,
O.Fedorov,
F.Bracher,
A.Chaikuad,
S.Knapp.
Dfg-1 Residue Controls Inhibitor Binding Mode and Affinity, Providing A Basis For Rational Design of Kinase Inhibitor Selectivity. J.Med.Chem. V. 63 10224 2020.
ISSN: ISSN 0022-2623
PubMed: 32787076
DOI: 10.1021/ACS.JMEDCHEM.0C00898
Page generated: Sat Aug 9 12:50:15 2025
ISSN: ISSN 0022-2623
PubMed: 32787076
DOI: 10.1021/ACS.JMEDCHEM.0C00898
Last articles
Mg in 3G5AMg in 3G5S
Mg in 3G58
Mg in 3G4T
Mg in 3G4L
Mg in 3G4K
Mg in 3G4I
Mg in 3G4G
Mg in 3G4F
Mg in 3G45