Potassium »
PDB 6dxz-6f3n »
6eo1 »
Potassium in PDB 6eo1: The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined)
Potassium Binding Sites:
The binding sites of Potassium atom in the The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined)
(pdb code 6eo1). This binding sites where shown within
5.0 Angstroms radius around Potassium atom.
In total 2 binding sites of Potassium where determined in the The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined), PDB code: 6eo1:
Jump to Potassium binding site number: 1; 2;
In total 2 binding sites of Potassium where determined in the The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined), PDB code: 6eo1:
Jump to Potassium binding site number: 1; 2;
Potassium binding site 1 out of 2 in 6eo1
Go back to
Potassium binding site 1 out
of 2 in the The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined)
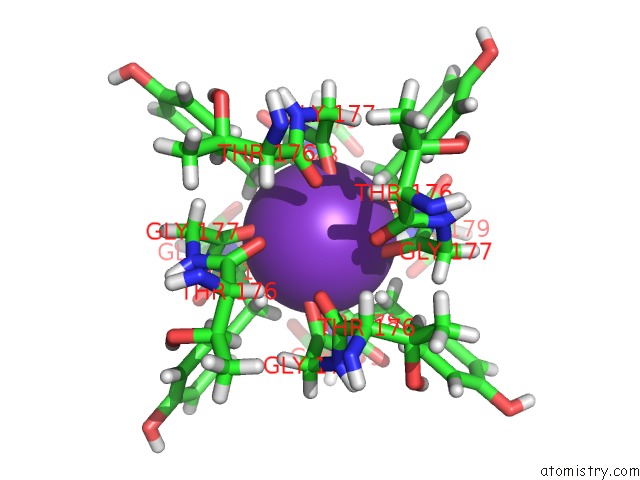
Mono view
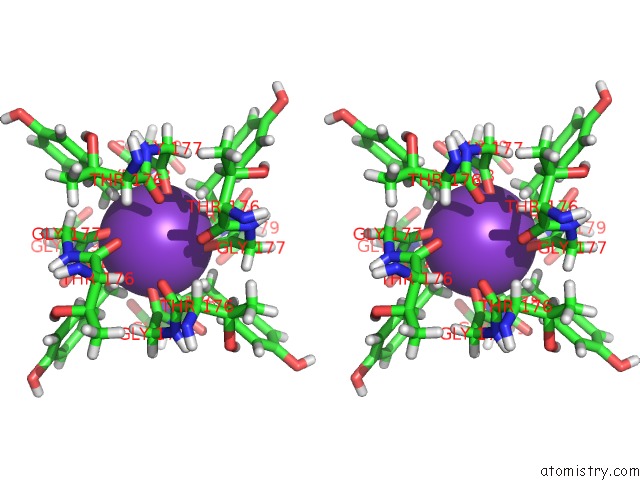
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 1 of The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined) within 5.0Å range:
|
Potassium binding site 2 out of 2 in 6eo1
Go back to
Potassium binding site 2 out
of 2 in the The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined)
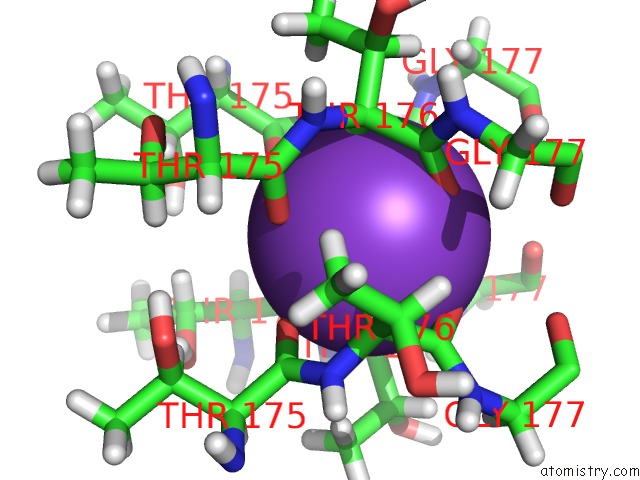
Mono view
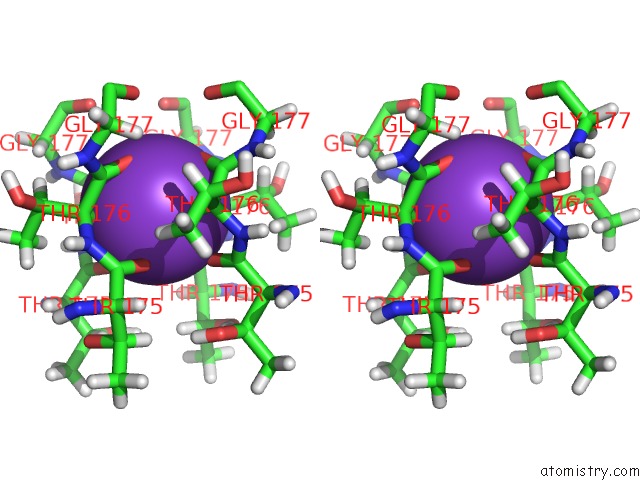
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 2 of The Electron Crystallography Structure of the Camp-Bound Potassium Channel MLOK1 (Pco-Refined) within 5.0Å range:
|
Reference:
J.Kowal,
N.Biyani,
M.Chami,
S.Scherer,
A.J.Rzepiela,
P.Baumgartner,
V.Upadhyay,
C.M.Nimigean,
H.Stahlberg.
High-Resolution Cryoelectron Microscopy Structure of the Cyclic Nucleotide-Modulated Potassium Channel MLOK1 in A Lipid Bilayer. Structure V. 26 20 2018.
ISSN: ISSN 1878-4186
PubMed: 29249605
DOI: 10.1016/J.STR.2017.11.012
Page generated: Mon Aug 12 16:00:20 2024
ISSN: ISSN 1878-4186
PubMed: 29249605
DOI: 10.1016/J.STR.2017.11.012
Last articles
Zn in 9J0NZn in 9J0O
Zn in 9J0P
Zn in 9FJX
Zn in 9EKB
Zn in 9C0F
Zn in 9CAH
Zn in 9CH0
Zn in 9CH3
Zn in 9CH1