Potassium »
PDB 4zun-5avw »
4zuq »
Potassium in PDB 4zuq: Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor
Protein crystallography data
The structure of Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor, PDB code: 4zuq
was solved by
C.Decroos,
D.W.Christianson,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 41.80 / 1.22 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 44.917, 120.511, 64.408, 90.00, 96.89, 90.00 |
| R / Rfree (%) | 13.7 / 15.2 |
Other elements in 4zuq:
The structure of Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor also contains other interesting chemical elements:
| Zinc | (Zn) | 4 atoms |
| Sodium | (Na) | 2 atoms |
Potassium Binding Sites:
The binding sites of Potassium atom in the Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor
(pdb code 4zuq). This binding sites where shown within
5.0 Angstroms radius around Potassium atom.
In total 2 binding sites of Potassium where determined in the Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor, PDB code: 4zuq:
Jump to Potassium binding site number: 1; 2;
In total 2 binding sites of Potassium where determined in the Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor, PDB code: 4zuq:
Jump to Potassium binding site number: 1; 2;
Potassium binding site 1 out of 2 in 4zuq
Go back to
Potassium binding site 1 out
of 2 in the Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor
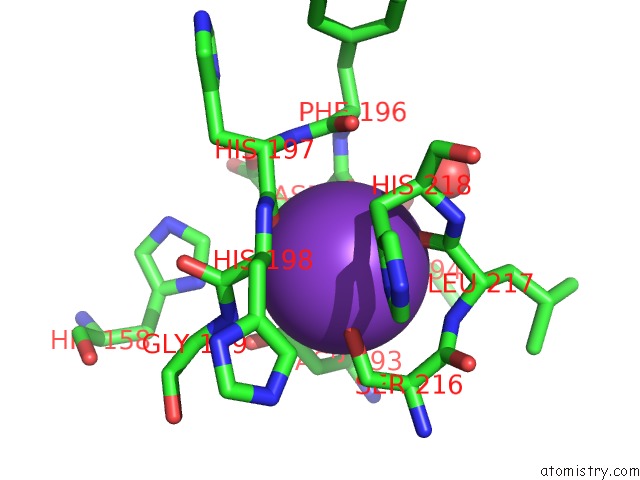
Mono view
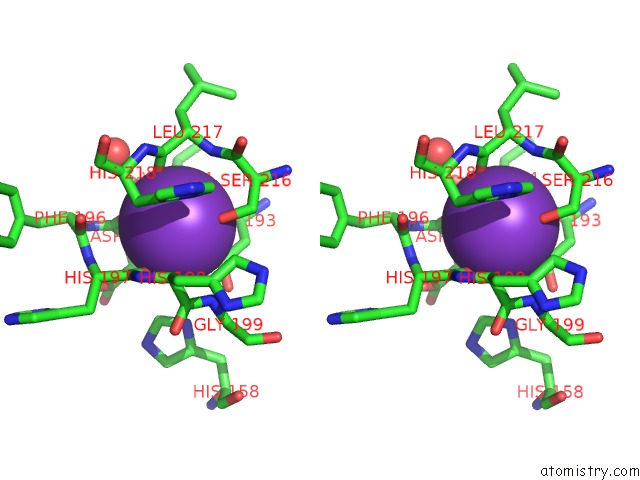
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 1 of Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor within 5.0Å range:
|
Potassium binding site 2 out of 2 in 4zuq
Go back to
Potassium binding site 2 out
of 2 in the Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor
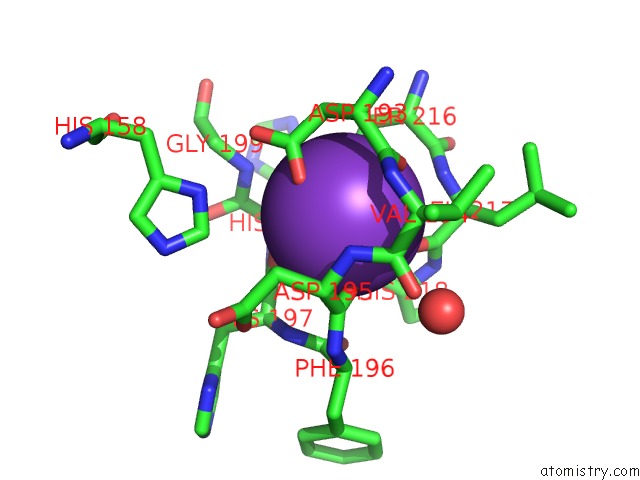
Mono view
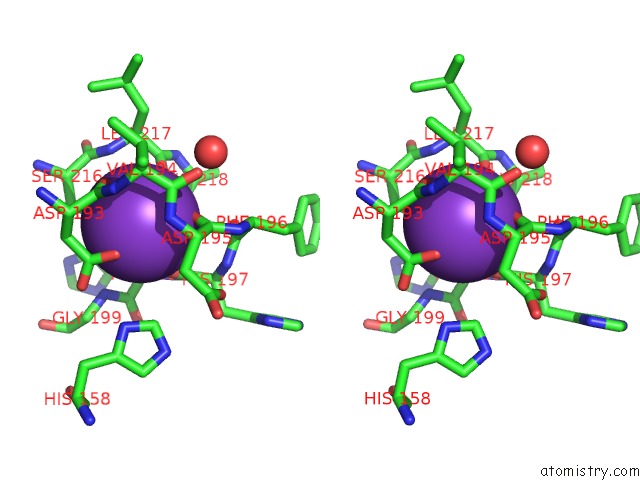
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 2 of Crystal Structure of Acetylpolyamine Amidohydrolase From Mycoplana Ramosa in Complex with A Hydroxamate Inhibitor within 5.0Å range:
|
Reference:
C.Decroos,
D.W.Christianson.
Design, Synthesis, and Evaluation of Polyamine Deacetylase Inhibitors, and High-Resolution Crystal Structures of Their Complexes with Acetylpolyamine Amidohydrolase. Biochemistry V. 54 4692 2015.
ISSN: ISSN 0006-2960
PubMed: 26200446
DOI: 10.1021/ACS.BIOCHEM.5B00536
Page generated: Sat Aug 9 08:25:44 2025
ISSN: ISSN 0006-2960
PubMed: 26200446
DOI: 10.1021/ACS.BIOCHEM.5B00536
Last articles
K in 8CFGK in 8CFE
K in 8CFD
K in 8CF8
K in 8CFC
K in 8CFB
K in 8CF1
K in 8CEP
K in 8CAI
K in 8CAZ