Potassium in PDB 8q0q: Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm
Enzymatic activity of Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm
All present enzymatic activity of Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm:
1.6.5.3; 1.6.99.3; 7.1.1.2;
1.6.5.3; 1.6.99.3; 7.1.1.2;
Other elements in 8q0q:
The structure of Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm also contains other interesting chemical elements:
| Magnesium | (Mg) | 1 atom |
| Iron | (Fe) | 28 atoms |
| Zinc | (Zn) | 1 atom |
Potassium Binding Sites:
The binding sites of Potassium atom in the Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm
(pdb code 8q0q). This binding sites where shown within
5.0 Angstroms radius around Potassium atom.
In total only one binding site of Potassium was determined in the Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm, PDB code: 8q0q:
In total only one binding site of Potassium was determined in the Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm, PDB code: 8q0q:
Potassium binding site 1 out of 1 in 8q0q
Go back to
Potassium binding site 1 out
of 1 in the Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm
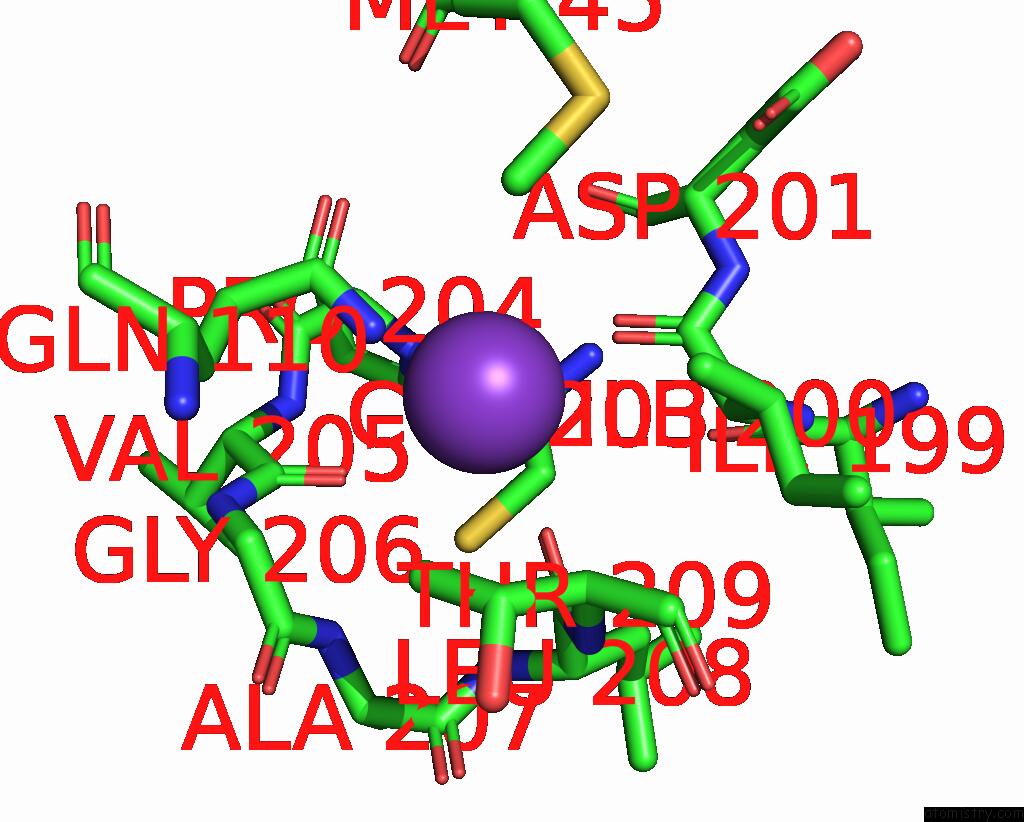
Mono view
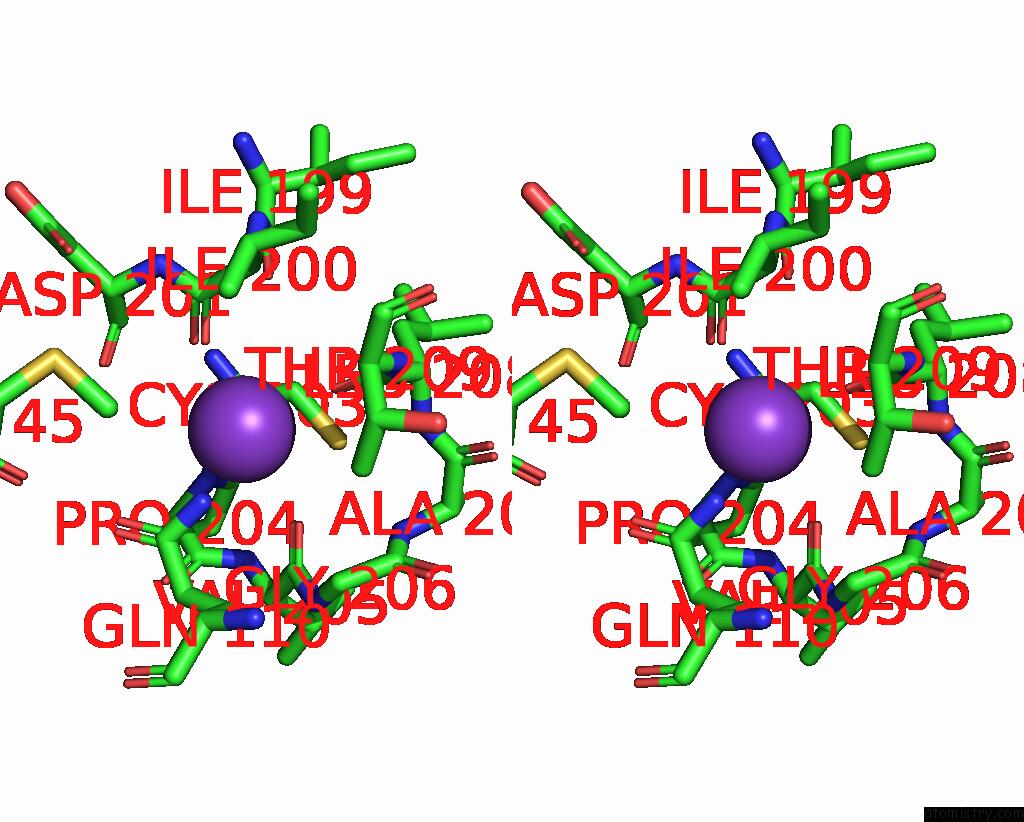
Stereo pair view

Mono view

Stereo pair view
A full contact list of Potassium with other atoms in the K binding
site number 1 of Outward-Facing, Slack Proteoliposome Complex I at 3.6 A. Initially Purified in Ddm within 5.0Å range:
|
Reference:
D.N.Grba,
J.J.Wright,
W.Fisher,
Z.Yin,
J.Hirst.
Molecular Mechanism of the Ischemia-Induced Regulatory Switch in Mammalian Complex I Science 2024.
ISSN: ESSN 1095-9203
DOI: 10.1126/SCIENCE.ADO2075
Page generated: Tue Aug 13 00:28:04 2024
ISSN: ESSN 1095-9203
DOI: 10.1126/SCIENCE.ADO2075
Last articles
Zn in 9MJ5Zn in 9HNW
Zn in 9G0L
Zn in 9FNE
Zn in 9DZN
Zn in 9E0I
Zn in 9D32
Zn in 9DAK
Zn in 8ZXC
Zn in 8ZUF